CE-IVD marked

Reproducible

Clinically validated

Short turnaround
Routine microbiome test
GA-map® Dysbiosis Test is a CE-IVD gut microbiome test suitable for routine testing. The assay is clinically validated and standardized, providing highly reliable and easy to interpret results. Easy assay workflow, high throughput capacity and integrated software enable short turnaround and automatic patient report generation.
NEW! AI-assisted interpretation with GA-map® Dysbiosis Test AInsight
AInsight is an AI-assisted interpretation platform designed specifically for GA-map® Dysbiosis Test results. The platform translates complex microbiome findings into structured insights to support clinician review and clearer patient discussions. AInsight retrieves scientific data exclusively from GA's curated Bacteria Compendium.
Decision-support only. Not a substitute for clinical judgment.
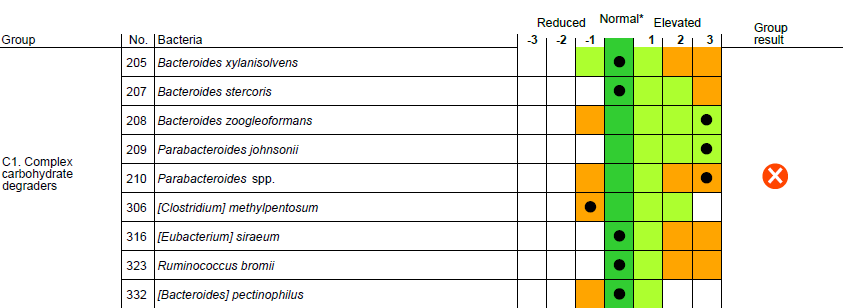
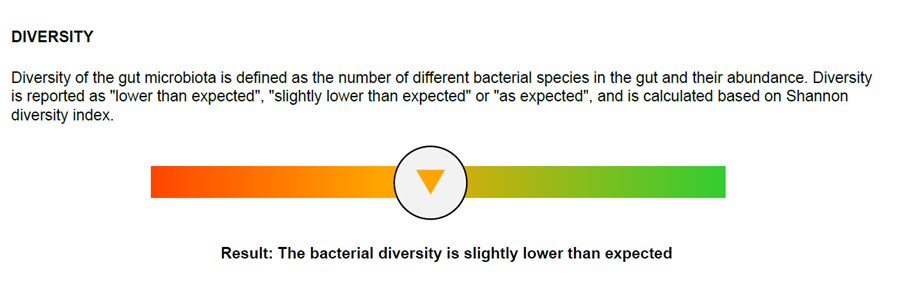
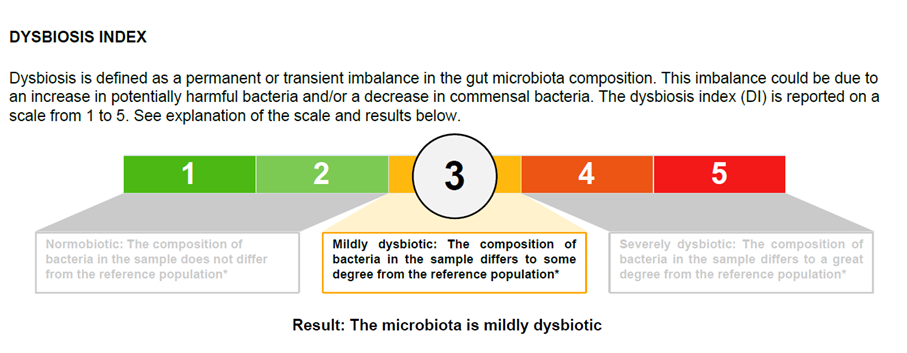
The clinically validated Dysbiosis Index (DI) is measured on a 5-point scale. DI value indicates the extent to which the bacteria profile of a sample deviates from a normal, healthy reference population.

Download Sample Report
Sample GA-map® Dysbiosis Test report form with explanatory appendix.
GA-map® Dysbiosis Test Sample Report Form
Unique algorithm
With GA-map® Dysbiosis Test, the need for comprehensive result calculations in gut microbiota testing is obsolete. The GA-map® Analyzer software processes the raw analysis data through a unique algorithm, which compares each sample to a healthy reference population to generate clinically meaningful results.
GA-map® Dysbiosis Test - assay specifications
Sample type
Stool
Read-out platform
MAGPIX®, Luminex®200™
Assay time
8 hours, pre-extracted samples
Output
Patient report including Dysbiosis Index, analyte abundance and microbiota functional profile
Assay format
48-plex, 1 well/sample

50+ publications
GA-map® Dysbiosis Test has been widely applied in clinical research, providing reliable microbiome data in the fields of gastroenterology and metabolic disorders.
Clinical validation of the GA-map® Dysbiosis Test and the specifics of the technology platform enable incorporating microbiome as a standardized parameter into clinical studies.
GA-map® Dysbiosis Test
Reference values
Relative abundance compared to healthy normal
No data processing steps
Patented algorithm providing ready-to-use results
Dysbiosis Index
Validated dysbiosis indicator
CE-IVD marked
Meets high safety, health, and environmental requirements
Result interpretation support
Access to AI-assisted result interpretation platform

We used GA-map® in our clinical research within IBS. We have great experience with GA-map® as a tool to select, monitor and follow-up patients.
Professor Magdy El-Salhy
Stord Hospital, Helse Bergen and the University of Bergen, Norway


There is still so much to learn about the microbiome, we are only just beginning to discover its importance, and the GA-map Test will help us do just that.
Emeritus Professor Peter Malfertheiner
Senior Professor at the Ludwig Maximillian University, University Clinic in Munich, Germany



